
Revolutionizing Protein Structure Analysis: DeepAFM Harnesses AI and Molecular Simulations
Exploring the Future of Protein Structure Analysis with DeepAFM
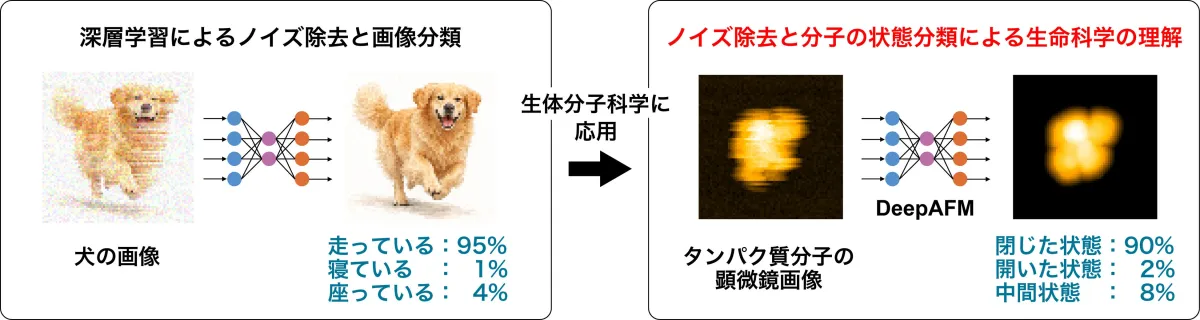
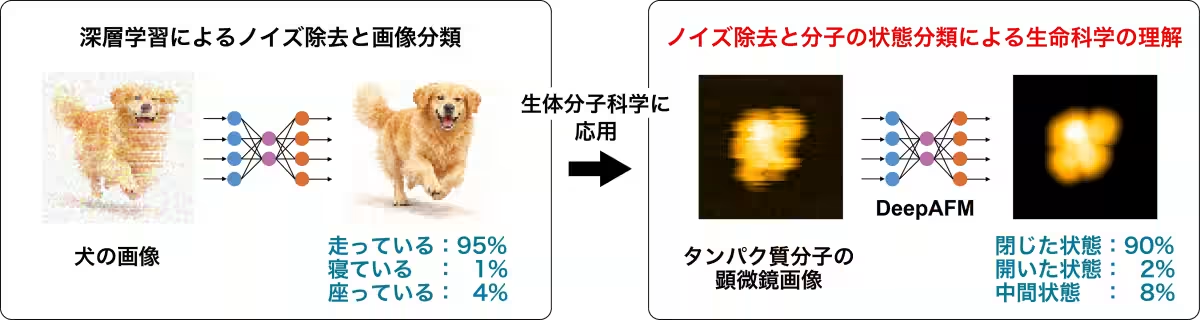
Scientific advancements often lead to breakthroughs that can significantly reshape our understanding of biological systems. One exciting recent development comes from Takaharu Mori, an associate professor at Tokyo University of Science, and his team, which has unveiled a revolutionary deep learning model known as DeepAFM. This innovative approach combines deep learning with molecular simulations to analyze high-speed atomic force microscopy (HS-AFM) images and accurately estimate protein structural states.
Understanding DeepAFM
DeepAFM stands as a powerful tool in the realm of structural biology. While HS-AFM has distinguished itself with the ability to monitor protein dynamics in real-time at the molecular level, it faces the challenge of noisy and distorted images that can complicate structural interpretation. DeepAFM tackles this by employing a wealth of artificial data generated through molecular dynamics simulations alongside deep learning techniques.
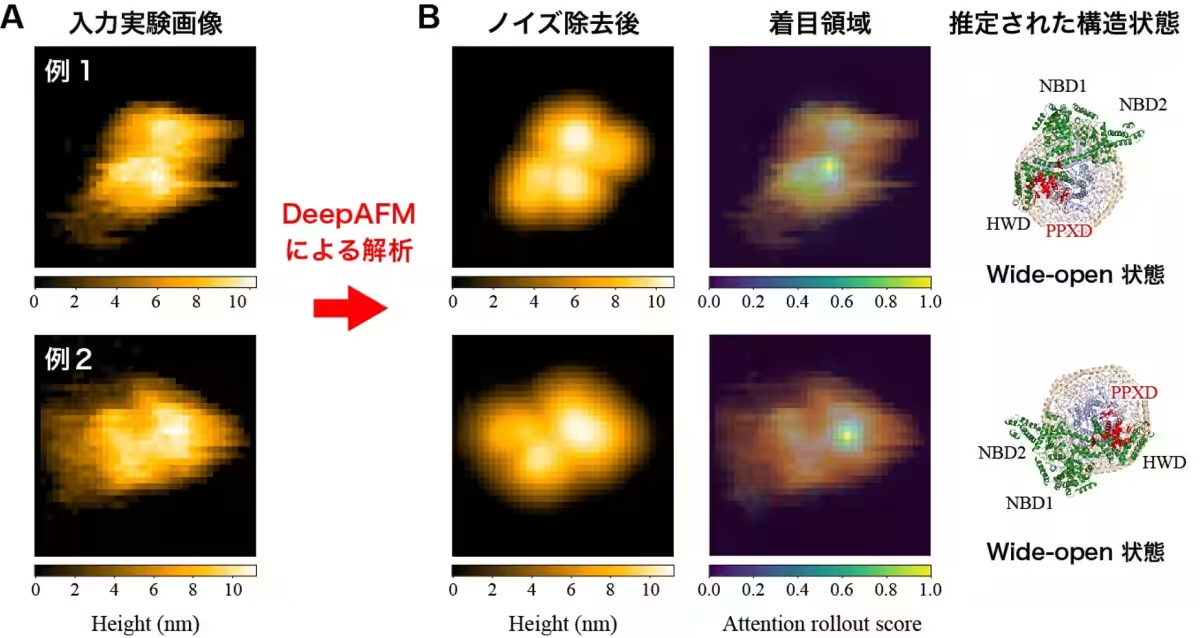
The output? A sophisticated method capable of efficiently classifying protein states and removing noise from experimental images. Through the application of DeepAFM, the team demonstrated that they could effectively deduce the structural states of proteins even when faced with the inherent difficulties posed by low-resolution microscopy images.
The Research Team's Journey
Joining forces with researchers from Nagoya University and Nara Institute of Science and Technology, Mori's group leveraged their expertise to create a novel image processing technique. This method brings unprecedented accuracy and efficiency to protein analysis, highlighting how creative combinations of computational and information sciences can propel our understanding of life sciences.
Traditional methods of protein state estimation relied on rigid fitting that often fell short in noisy conditions. By integrating mechanisms like molecular dynamics and state-of-the-art deep learning algorithms, the researchers have established a new standard for image interpretation in biomolecular studies.
Research Outcomes and Potential Applications
The findings from this study were published in the Journal of Chemical Information and Modeling, indicating a major step forward in biophysical analysis. The applications for DeepAFM could be vast, reaching across various biological systems. It opens the door for advancements that not only enhance our understanding of molecular interactions but also help in the design of new therapeutic strategies.
For proteins that undergo significant conformational changes during functional activation, precise structural analysis is crucial. The capability of DeepAFM to discern fine details of protein motion offers new opportunities to study enzymatic reactions, signaling pathways, and transport mechanisms at an unprecedented resolution.
Future Prospects
Mori emphasizes the importance of continued exploration in this area, suggesting that innovations in artificial intelligence could further enrich human analytical capabilities. As research progresses, DeepAFM is positioned to enhance the precision of protein characterization methods, providing critical insights into the biological mechanisms that underpin health and disease.
In conclusion, the introduction of DeepAFM represents a noteworthy stride in the intersection of deep learning and molecular biology. It promises not only to advance the accurate analysis of protein structures but also to contribute to a deeper understanding of life's fundamental processes. As we look to the future, this pioneering approach may well change the course of research in the life sciences.
Topics Health)









【About Using Articles】
You can freely use the title and article content by linking to the page where the article is posted.
※ Images cannot be used.
【About Links】
Links are free to use.